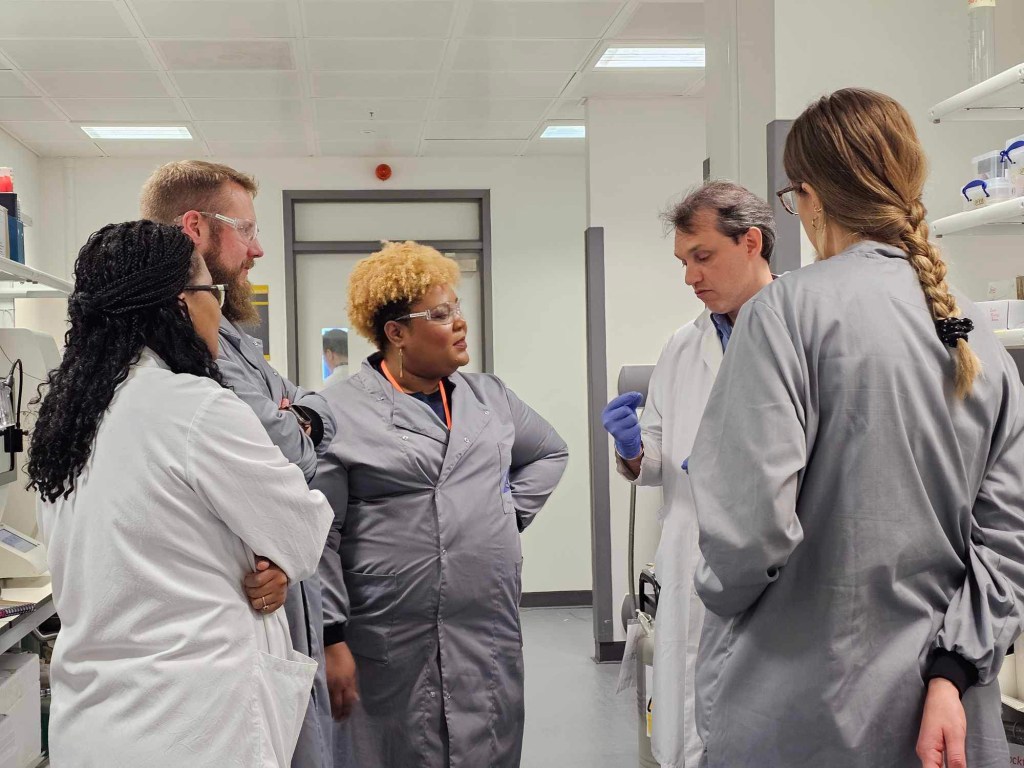
The Hinton Laboratory is a Molecular Physiology and Biophysics research program dedicated to defining how ATF4-dependent stress signaling regulates organelle structure, morphology, and inter-organelle communication across health, aging, and disease. Our work is grounded in the principle that mitochondrial shape is dynamic, programmable, and functionally instructive. We believe mitochondrial morphology is not passive. Shape is defined by underlying membrane architecture, transcriptional control, and organelle interaction networks, and this structure directly determines metabolic and signaling function.
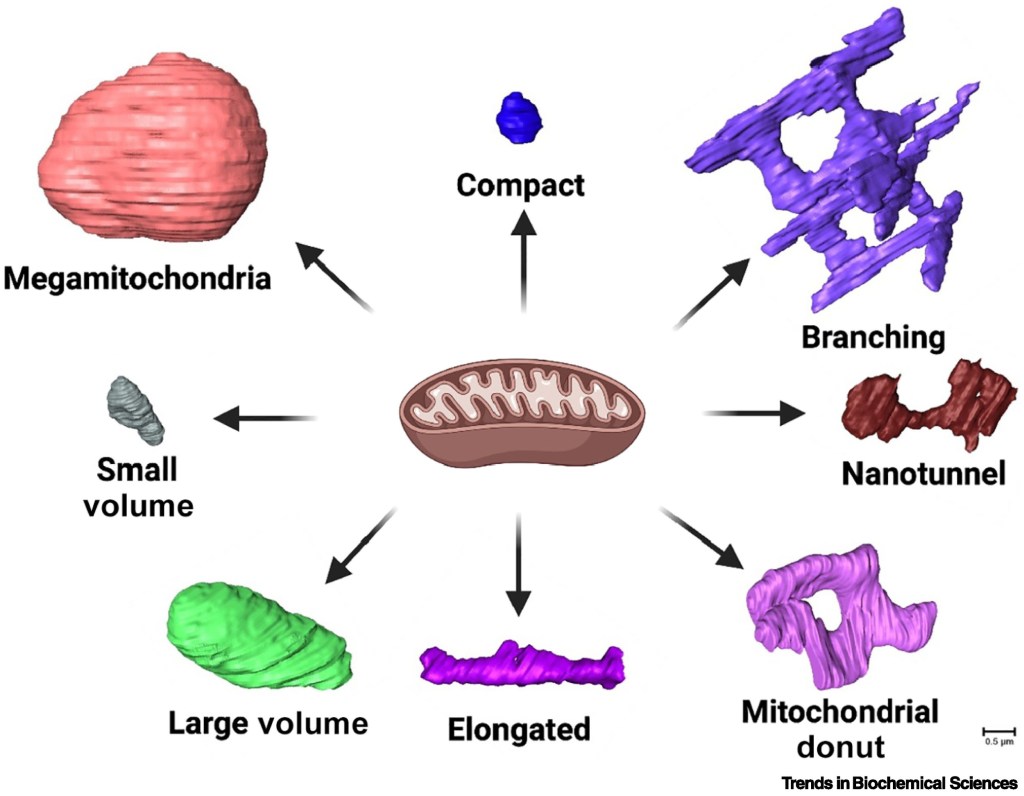
Our laboratory operates under the overarching hypothesis that stress-responsive transcriptional programs, particularly ATF4, dynamically remodel mitochondrial membrane architecture and organelle contact networks, thereby programming mitochondrial shape states that determine bioenergetic performance, redox balance, and cellular adaptation. We propose that mitochondrial structural states, including megamitochondria, compact forms, branching networks, nanotunnels, elongated structures, donut-shaped mitochondria, and fragmented small-volume forms, represent adaptive morphological programs encoded by stress signaling and metabolic demand. In this framework, structure defines shape, shape encodes function, and function determines physiological outcome.
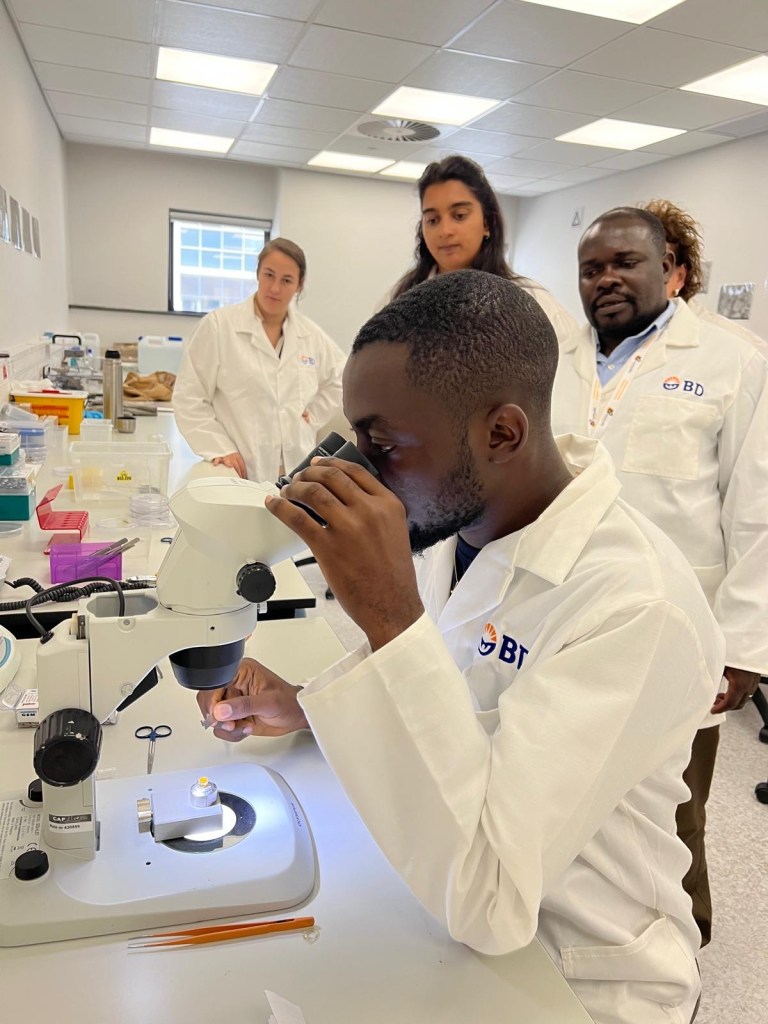

At the center of our research is ATF4, a master regulator of the Integrated Stress Response. We investigate how ATF4 coordinates mitochondrial dynamics, cristae remodeling, mitochondrial DNA nucleoid organization, lysosomal regulation, lipid droplet–mitochondria interactions, and endoplasmic reticulum–mitochondria contact site architecture to control cellular energetics, proteostasis, redox balance, and metabolic adaptation. Through these mechanisms, we define how transcriptional stress signaling reshapes organelle networks in skeletal muscle, cardiometabolic disease, hypertension, aging, and systemic stress-related disorders.onceptually, our laboratory advances a unifying framework we term Organelle Plasticity Biology, the principle that organelle architecture is dynamically programmed by stress-responsive transcriptional and metabolic networks. We are active contributors to and leaders within the mitochondrial biology field, the organelle contact site field, the stress biology field, the electron microscopy field, and the broader microscopy and quantitative imaging sciences. A defining feature of the laboratory is our emphasis on spatial and structural cell biology. Using Serial Block-Face Scanning Electron Microscopy, Focused Ion Beam Scanning Electron Microscopy, Correlative Light and Electron Microscopy, confocal imaging, volumetric reconstruction, and quantitative morphometric analysis, we define the structural principles governing mitochondrial membrane curvature, cristae density, network topology, nucleoid positioning, and organelle spatial organization within metabolically active tissues.
We are not only practitioners of electron microscopy but active innovators in the field. The laboratory develops next-generation electron microscopy workflows, volumetric reconstruction strategies, and artificial intelligence–driven segmentation pipelines to classify mitochondrial ultrastructure, membrane topology, and organelle interaction networks across aging, metabolic disease, endoplasmic reticulum stress, and mechanical stimulation paradigms. Through transcriptomic profiling, chromatin occupancy analysis, Seahorse bioenergetic flux measurements, and computational modeling, we map transcriptional programs that couple mitochondrial bioenergetics, mitochondrial DNA regulation, lysosomal pathways, redox homeostasis, and adaptive stress responses. This integrated framework positions ATF4 as a central regulator linking transcription, membrane architecture, and physiological function.
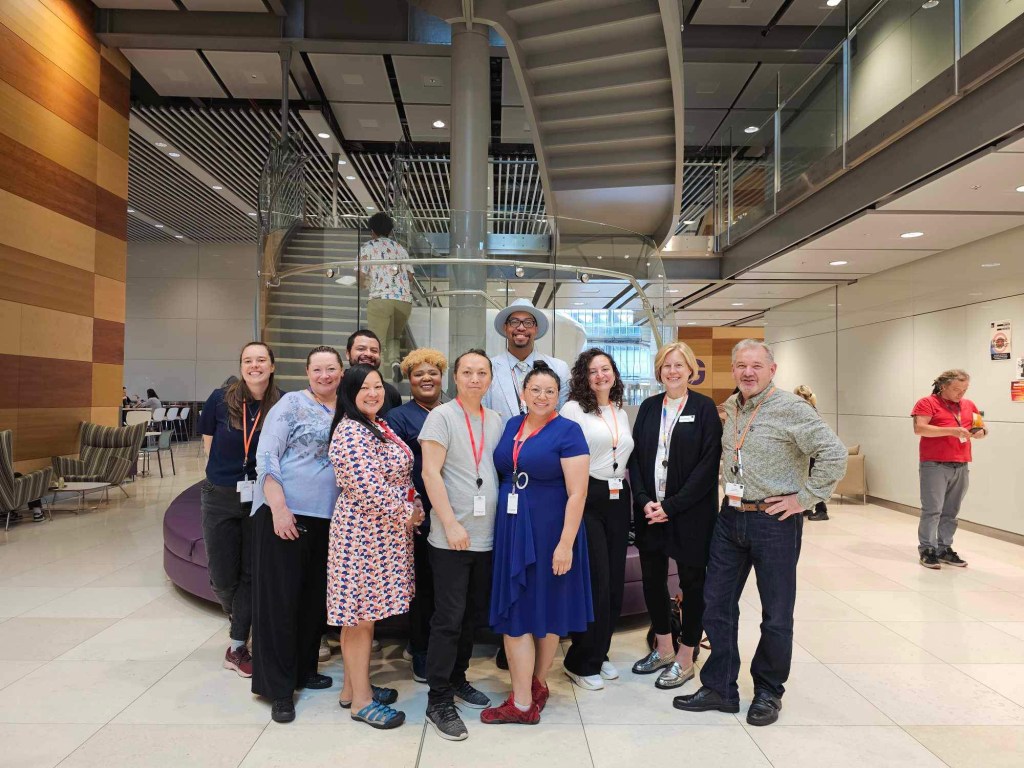

Our long-term goal is to redefine therapeutic strategy by demonstrating that mitochondrial structure is programmable and that restoring adaptive shape states may restore metabolic resilience. By bridging ultrastructure with transcriptional control and systems physiology, we aim to uncover new avenues for intervention in cardiometabolic disease, skeletal muscle dysfunction, and aging-related disorders. Beyond discovery, the Hinton Laboratory is deeply committed to training and mentorship. We cultivate hybrid scientists fluent in imaging, molecular genetics, physiology, and computational biology, and we provide structured mentorship, interdisciplinary exposure, leadership development, and quantitative rigor to prepare trainees to lead at the intersection of mitochondrial biology, organelle contact science, and advanced microscopy. We believe that advancing the mitochondrial and organelle biology fields requires both scientific innovation and intentional development of diverse scientific talent.